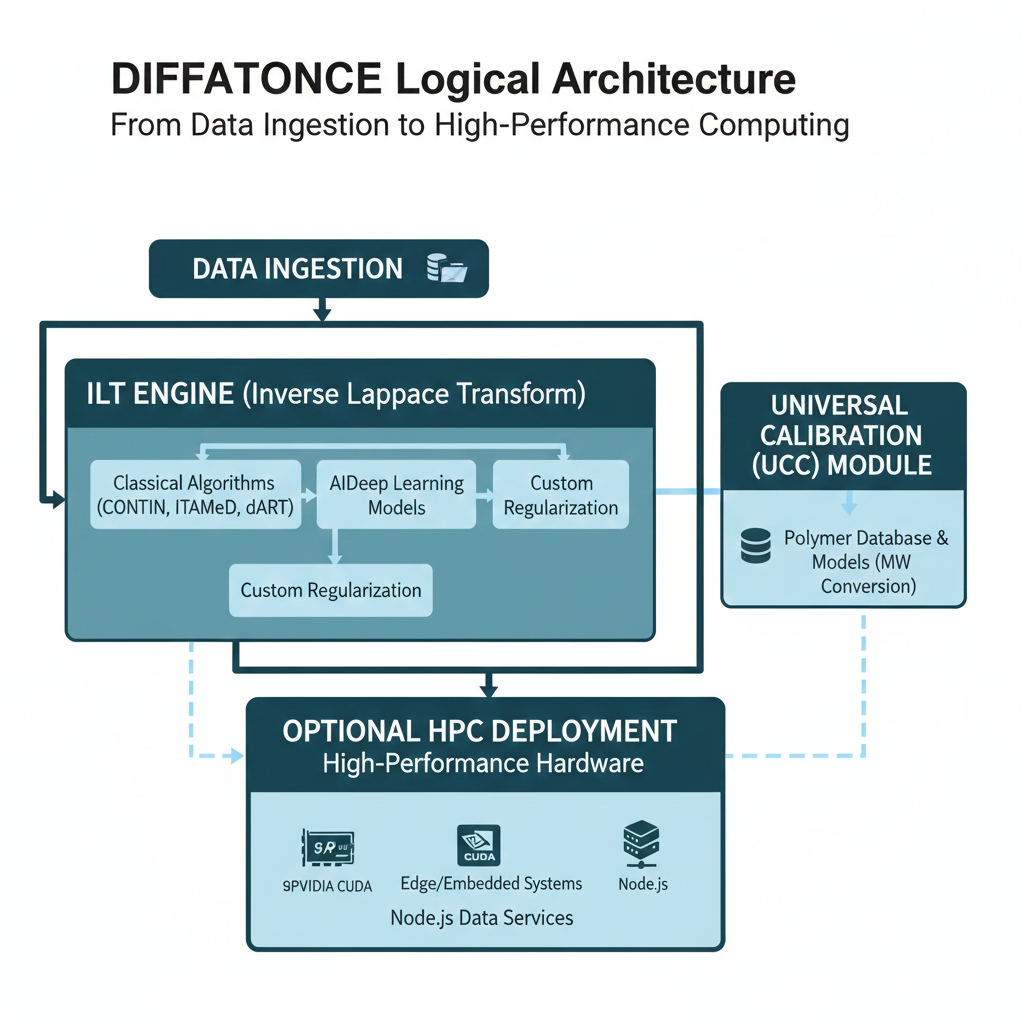
Universal calibration (UCC) and molecular weight
Once diffusion coefficient distributions have been obtained, DiffAtOnce applies
universal calibration (UCC) models developed by the NMRMBC group to transform
D into molecular weight distributions, minimizing the dependence on solvent viscosity and other
experimental conditions.
The models incorporate the β parameter and the fractal dimension of the polymer, in line with
Flory's theory and empirical relationships between hydrodynamic radius, diffusion coefficient
and molecular weight. The software accesses a database of UCC curves (for polystyrene,
polypropylene, globular proteins, etc.) and allows extensions with new calibrations generated
within FUNPOLYMER.
UCC parameters are stored in a database server and exposed through dynamic libraries, so that
other applications can reuse the calibration curves for predicting molecular weight from D,
viscosity and polymer type.
Extended diffusion NMR (ediffNMR)
Beyond classical diffusion experiments, DiffAtOnce includes the
extended diffusion NMR (ediffNMR) methodology, in which both gradient duration
and gradient strength are varied. This ensures that the data set is informative for both low and
high molecular weight species, while keeping experimental time under control.
This methodology requires a full adaptation of ILT algorithms to bi-multiexponential decays
depending on δ and G, and has been developed within FUNPOLYMER together with the optimization
of pulse sequences, noise assessment and resolution of components with very close diffusion
coefficients.